Archive
Gene network reconstruction from transcriptional dynamics
D. R. Bickel, Z. Montazeri, P.-C. Hsieh, M. Beatty, S. J. Lawit, and N. J. Bate, “Gene network reconstruction from transcriptional dynamics under kinetic model uncertainty: A case for the second derivative,” Bioinformatics 25, 772-779 (2009).
Fold change estimation versus hypothesis testing
D. R. Bickel and C. M. Yanofsky, “Validation of differential gene expression algorithms: Application comparing fold change estimation to hypothesis testing,” Preprint, Ottawa Institute of Systems Biology, COBRA Preprint Series, Article 50, available at tinyurl.com/bw9jod (2009).
Strength of evidence for composite hypotheses
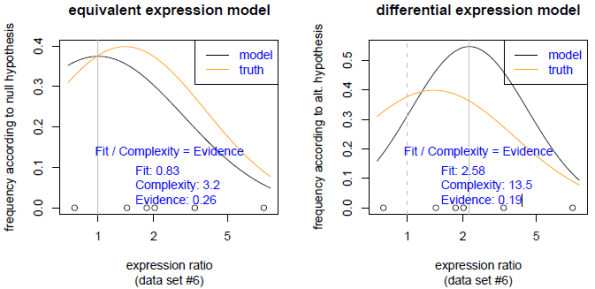
D. R. Bickel, “The strength of statistical evidence for composite hypotheses with an application to multiple comparisons,” Preprint, Ottawa Institute of Systems Biology, COBRA Preprint Series, Article 49, available at tinyurl.com/7yaysp (2008).
Preprint servers for statistical bioinformatics
The COBRA Preprint Series was selected for the dissemination of the Statomics Lab’s working papers because it offers more flexibility than the other main services for statistical genomics preprints:
- Unlike arXiv, COBRA accepts PDF files generated by LaTeX. Preparing LaTeX source code specifically for a preprint server can require large time investments.
- Unlike Nature Precedings, COBRA does not require authors to irrevocably license their work for commercial reuse. Such licenses might conflict with the interests of some commercial publishers since they allow competitors to distribute the preprints for profit.
Local false discovery rate software
Zahra Montazeri and David Bickel developed empiricalBayes, an R software bundle that provides a simple solution to the extreme multiple testing problem. It contains two packages:
- localFDR estimates local false discovery rates given a vector of p-values.
- HighProbability determines which p-values are low enough that their alternative hypotheses can be considered highly probable.
Application: cis- & trans-effects on gene expression
M. Guo, S. Yang, M. Rupe, B. Hu, D. R. Bickel, L. Arthur, and O. Smith, “Genome-wide allele-specific expression analysis using Massively Parallel Signature Sequencing (MPSS) reveals cis- and trans-effects on gene expression in maize hybrid meristem tissue,” Plant Molecular Biology 66, 551-563 (2008).
Estimating levels of differential gene expression
D. R. Bickel, “Correcting the estimated level of differential expression for gene selection bias: Application to a microarray study,” Statistical Applications in Genetics and Molecular Biology 7 (1) 10, http://www.bepress.com/sagmb/vol7/iss1/art10 (2008).
Mode estimation
Paul Poncet’s modeest package implements the half-range mode, the half-sample mode, and the mode-based skewness of D. R. Bickel, “Robust estimators of the mode and skewness of continuous data,” Computational Statistics and Data Analysis 39, 153-163 (2002).
You must be logged in to post a comment.